Staff Profile of Dr Martin Llewellyn

- Name
- Dr Martin Llewellyn
- Position
- Research Fellow
- Martin.Llewellyn@glasgow.ac.uk
- Phone
- Skype: martin_llewellyn
- Location
- Genomics and Conservation of Aquatic Species, Department of Biology, Universite de Laval, Quebec G1V 0A6, Canada
and
School of Natural Sciences, Bangor University, Deiniol Road, Bangor, Gwynedd, LL57 2UW, UK
About
Molecular parasitology: parasites are abundant and successful organisms distributed throughout the tree of life. A parasite’s genetic diversity provides valuable clues to track the its dispersal and transmission. Population genetic (and increasingly genomic) tools can thus provide insights for epidemiologists and health / veterinary professionals.
Complex infections: parasitic organisms, rarely, if ever, exist as genetically homogeneous infections within their host. Multiple genetic lineages of the same parasite species often infect the same host at the same time, with consequences for immune evasion, transmission scenarios and drug resistance.
The accessory microbiome: Most parasitic infections occur among a soup of billions of other non-host cells - the commensal microbiome. Thus parasite genetic lineages not only interact with each other and the host during parasitic infections, but also with microbiota present on/in the host. Early indications suggest that accessory microbiota have an important role in defining disease outcomes.
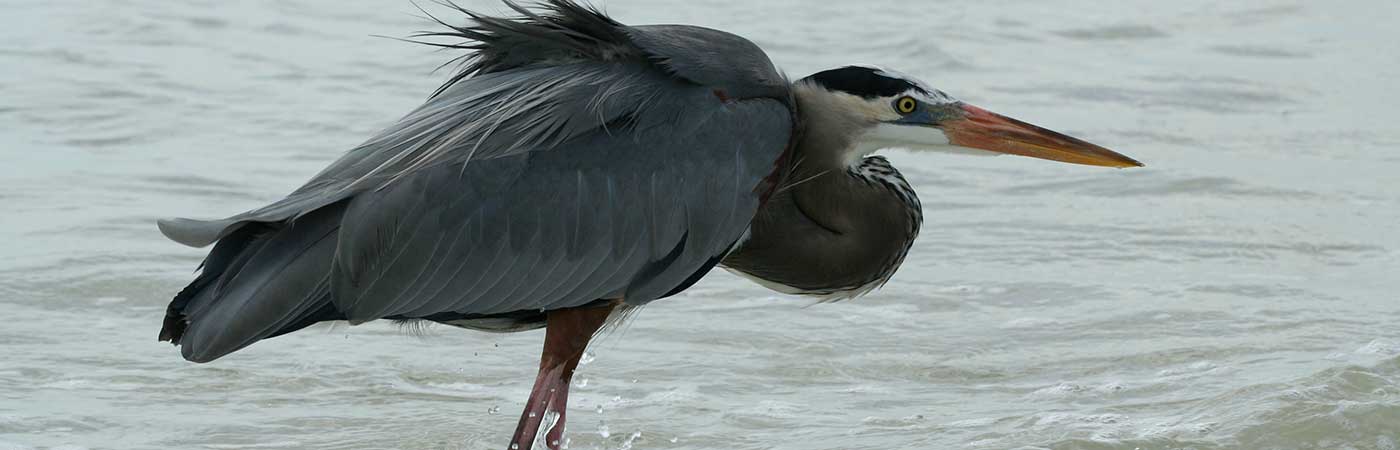
My research explores these themes in multiple parasitic systems: trypanosomes and their mammalian hosts (Trypanosoma cruzi and Leishmania infection) and the copepod pathogen Lepeophtheirus salmonis and its salmonid hosts.
.
CV
Education
2009 PhD, London School of Hygiene and Tropical Medicine – “The molecular epidemiology of Trypanosoma cruzi infection in wild and domestic transmission cycles, with special emphasis on multilocus microsatellite analysis”.
2002 BA (Hons), Oxford University, Biological Sciences
Career
2012–2015 – Marie Curie Research Fellow, Department of Biology, Universite de Laval, Quebec & School of Biological Sciences, Bangor University, UK.
2009–2012 – ChagasEpiNet – Post-doctoral researcher – London School of Hygiene and Tropical Medicine.
2010 – Visiting Research Fellow – Facultad de Medicina, Universidad Mayor de San Simon, Cochabamba, Bolivia.
2007 – Visiting Research Fellow – Instituto de Medicina Tropical, Universidad Central De Venezuela, Caracas.
2005 & 2007 – Visiting Research Fellow – Centro Nacional de Enfermedades Tropicales, Santa Cruz, Bolivia
2004 – Visiting Research Fellow – Instituto Evandro Chagas, Belem, Para, Brazil.
Grants and Awards
2012–2015 – FishProBio Marie Curie Fellowship – Salmonid microbiomes in health and disease, Principal Investigator – €248,220
2011–2013 – DeepCruzi transnational access project, ESGI consortium – Intra-host genetic diversity in T. cruzi, Principal Investigator – €85,000
2009–2012 – ChagasEpiNet EC Fp7 Consortium – Molecular Epidemiology of Chagas disease, named researcher – €2,876,507
2009 – De Lazslo Foundation – Chagas disease origins in man – £4,000
2007 – Gordon Smith Scholarship – Emergent Chagas Disease in Northern Bolivia – £5,000
2005 – Swire Trust Travelling Scholoarship – The molecular epidemiology of human trypansomiasis in Venezuela – £5,000
Research
Salmo salar microbiome project 2013-2015
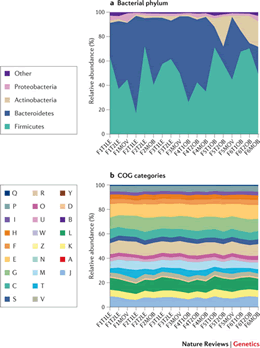 |
Figure 1 - Taxonomic diversity contrasts with functional stability between human microbiomes. COG categories refer to functionally similar genetic sequences. From Cho and Blaser, 2012, Nat Genetics. |
Atlantic salmon is the primary finfish aquaculture species in Canada, the UK, Scandinavia and Chile. The farmed S. salar market is worth over 7 billion USD annually. Nutrition is perhaps the major consideration in Salmonid aquaculture. Efficient nutrient transfer in terms of feed to weight ratios, as well as the provision of non-marine protein sources, are vital to mitigate the impact of salmon farming. Furthermore communicable disease has an increasing impact on farmed fish, with several emergent microbial pathogens in circulation.
Like the human gut microbiome project, an understanding of the S. salaar microbiome will provide insight into the role microbial species have in nutrient absorption and metabolism, immunity and disease. Unlike in humans, where much of the microbiome is transmitted to juveniles within families and social groups, S. salar must selectively recruit all their commensal bacteria from the external environment. We intend to explore common characteristics of these communities between fish in different ecological settings. To achieve this objective we propose to take a meta-genomic approach to identify common ‘services’ provided by the microbiome (see Figure 1). In this project, we propose to characterize the microbiome of adult Salmo salar across its wild and captive range including individuals from marine, estuarine, and freshwater environment.
The S. salar project is designed to act as a precursor to the study of the microbiome as accessory to parasitic disease.
Funded by Marie-Curie International Outgoing Fellowship, Martin Llewellyn
Sea lice and the mucosal microbiome 2013 - 2015
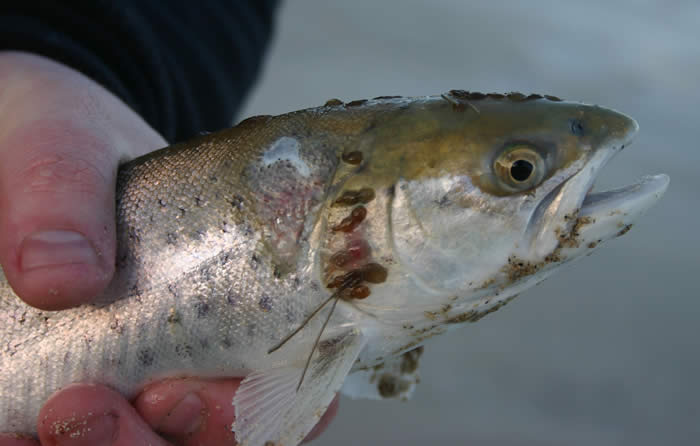 |
Figure 2 – Heavily infected S. salar sub adult with L. salmonis (www.robedwards.com) |
Sea lice, copepod fish ectoparasites, are a major burden of disease on commercially reared salmon. The annual cost of infection is considered to be in excess of €300 million. Several species are implicated, principally L. salmonis and members of genus Caligus. Sea lice feed on host mucus, skin and underlying tissue, causing legions that precede secondary bacterial infections.
In this project, we are to examining the interaction between L. salmonis infection and the host commensal microbiome throughout the L. salmonis lifecycle.
We aim to evaluate microbiome dynamics prior and during copepodid attachment, chalimus, pre-adult and adult development in wild and farmed fish. We are exploring explore infestation density effects, mucous composition (cortisol, immunochemistry), host heath status and the ecological succession of opportunistic pathogens that accompany lesion formation.
Funded by Marie-Curie International Outgoing Fellowship, Martin Llewellyn & ENGAGE project, NSERC, Canada, Nicolas Derome.
DeepCruzi – Deep sequencing to resolve cryptic Trypanosoma cruzi multiclonality in single hosts, 2011–2015
 |
Figure 3 - Heatmap showing amplicon deep sequencing profiles of Trypanosoma cruzi diversity within in individual hosts (Llewellyn et al 2014, in prep). |
As part of an FP7 Infrastructure consortium – European Sequencing and Genotyping Infrastructure (ESGI), we are applying amplicon and whole genome deep sequencing to understand T. cruzi diversity within the human individual hosts in a collaboration between (London School of Hygiene and Tropical Medicine, Centre National de Genotypage, Paris). The aim of this project is to evaluate T. cruzi intra-host diversity in the context of several epidemiological phenomena: congenital transmission, progression to symptomatic disease, oral disease transmission and parasite
adaptation to long-term host immune exposure.
Funded by EC FP7 transnational access grant from ESGI Consortium
Trypanosoma cruzi landscape genetics, Ecuador and Peru - 2014 - 2017
Neglected Tropical Diseases affect approximately 2.7 billion people worldwide and are
widely distributed, especially among the tropics. The dispersal and distribution of the
pathogens that cause NTDs are strongly influenced by the spatial complexity of their
physical and biotic environment. Thus, spatial analyses of abiotic and biotic variables can serve to accurately predict the emergence, occurrence, and spread of disease.

Figure 4 – Trypanosoma cruzi and the local vector species – Rhodnius ecuadoriensis
In this project we focus on Chagas disease in an area of disease transmission in Southern Ecuador and Northern Peru. Chagas disease is caused by the single-celled kinetoplastid Trypanosoma cruzi, and in our study region the local insect vector species is Rhodnius ecuadoriensis. Our aim is to establish the population genetic/genomic diversity of both vector and parasite across the study region by high resolution (genome scale) genotyping. To evaluate the interconnectivity of different vector and parasite populations, and to predict potential dispersal routes or barriers to geneflow, we adopt a landscape genetics approach.
Our aim is to define what environmental factors and what spatial scales are relevant in defining parasite and vector population distribution. Such descriptions of diversity are key to measuring and predicting dispersal. Our evaluation of barriers to gene flow will feed directly into a rationale for effective local disease control. Furthermore our project will provide a model with which to approach the study of the dispersal and distribution of other NTD agents.
Funded by the NIH in collaboration with Ohio University & University of Montana (USA) Pontificia Universidad Católica del Ecuador (PUCE) (Ecuador), Karolinska Institute (Sweden), London School of Hygiene and Tropical Medicine (London).
Leishmaniasis in the Brazilian Amazon: the role of accessory microbiota in disease progression, pathobiology and immunity. 2015-2017
Leishmania spp. infection represents a serious public health burden in Brazil, with 35,000 suspected cases annually. Disfiguring and occasionally fatal cutaneous and muco-cutaneous forms of Leishmaniasis are most common in the Amazon region of Brazil. Differential clinical severity and drug-resistance profiles are widely reported between patients. There is thus an urgent need to identify what factors might be responsible for different patient outcomes. Leishmania species and genotype have a role in defining disease severity, as does the host immune response. Less commonly considered is the role that secondary bacterial infections and commensal skin microbiota have in modulating pathology and immunity. In this multidisciplinary project we propose to use state-of-the-art techniques to monitor Leishmania lesion-associated microbial diversity and host immune response to reveal the factors that underlie cutaneous Leishmaniasis severity and progression in Amazonian Brazil.
Figure 1 - Mucocutaneous leishmaniasis (left) represents a disfiguring and potentially life-threatening progression from cutaneous leishmaniasis (right)
Funded by FAPERJ, Brazil and FAPERJ-MRC, UK (Newton-CONFAP). Collaborators Chris Quince (Warwick, UK), Elisa Cupilillo (FIOCRUZ, Brazil)
Recent Projects
ChagasEpiNet, a 15 partner EC FP7 consortium whose stated aim is to “elucidate the epidemiology of the genetic lineages of Trypanosoma cruzi, for improved understanding and prevention of Chagas disease”. 2009–2012
Publications
Llewellyn, MS Messenger, LA Luquetti A, Garcia L, Torrico F, Tavares, S Cheaib, B Derome, N Delepine, M Baulard, C Deluze, JF Sauer, S Miles, MA2015 Deep sequencing of the Trypanosoma cruzi GP63 surface proteases reveals diversity and diversifying selection among chronic and congenital Chagas disease patients. Submitted
Messenger, LA; Garcia, AL; Vanhove, M; Huaranca, C; Bustamante, M; Torrico, M; Torrico, F; Miles, MA; Llewellyn, MS 2015 Ecological host fitting of Trypanosoma cruzi TcI in Bolivia: mosaic population structure, hybridization and a role for humans in Andean parasite dispersal. Molecular Ecology. Accepted
Messenger LA, Yeo M, Lewis MD, Llewellyn MS, Miles MA. 2014 Molecular Genotyping of Trypanosoma cruzi for Lineage Assignment and Population Genetics. Methods Mol Biol. 1201:297-337.
Diosque P, Tomasini N, Lauthier JJ, Messenger LA, Monje Rumi MM, Ragone PG, Alberti-D'Amato AM, Pérez Brandán C, Barnabé C, Tibayrenc M, Lewis MD, Llewellyn MS, Miles MA, Yeo M. 2014 Optimized multilocus sequence typing (MLST) scheme for Trypanosoma cruzi. PLoS Negl Trop Dis. 8:e3117
Ramírez JD, Llewellyn MS. 2014 Reproductive clonality in protozoan pathogens - truth or artefact? Molecular Ecology 23: 4195–4202,
Baleela, R Llewellyn MS Fitzpatrick, S Kuhls, K Schönian, G Miles MA Mauricio IL 2014 Leishmania donovani populations in Eastern Sudan: temporal structuring and a link between human and canine transmission. Parasites and Vectors 7:496
Lima, V. Jansen, AM, Messenger, LM. Miles, MA. Llewellyn, MS. 2014 Wild Trypanosoma cruzi I genetic diversity in Brazil suggests admixture and disturbance in parasite populations from the Atlantic Forest region. Parasites and Vectors 7:263
Llewellyn MS, Boutin, SA. Hossein, S, Derome, N 2014 Teleost microbiomes: progress towards characterisation, manipulation and applications in aquaculture and fisheries. Frontiers in Microbiology. 5:207
Segovia M, Carrasco HJ, Martínez CE, Messenger LA, Nessi A, Londoño JC, Espinosa R, Martínez C, Alfredo M, Bonfante-Cabarcas R, Lewis MD, de Noya BA, Miles MA, Llewellyn MS. 2013 Molecular epidemiologic source tracking of orally transmitted Chagas disease, Venezuela. Emerg Infect Dis. 19:1098-101
Tomasini N, Lauthier JJ, Llewellyn MS, Diosque P. 2013 MLSTest: novel software for multi-locus sequence data analysis in eukaryotic organisms. Infect Genet Evol. 20:188-96.
Zumaya-Estrada FA, Messenger LA, Lopez-Ordonez T, Lewis, MD , Flores-Lopez CA, Martínez-Ibarra JA, Pennington PM, Cordon-Rosales C, Carrasco HJ, Segovia M, Miles MA, Llewellyn MS 2012 North American import? Charting the origins of an enigmatic Trypanosoma cruzi domestic genotype. Parasites and Vectors. 5:226 doi:10.1186/1756-3305-5-226
Franzén, O. Talavera-López, C. Ochaya, S. Butler, CE. Messenger, LA. Lewis, MD. Llewellyn, MS. Marinkelle, CJ Tyler, KM Miles MA, Andersson BA (2012) Comparative genomic analysis of human infective Trypanosoma cruzi lineages with the bat-restricted subspecies T. cruzi marinkellei. BMC Genomics. 13:531
Ramírez, JD, Guhl, F, Messenger, LA, Lewis, MD, Montilla, M, Cucunuba, Z, Miles, MA; Llewellyn, MS. (2012) Contemporary cryptic sexuality in Trypanosoma cruzi. Molecular Ecology. 21: 4216–4226,
Carrasco HJ, Segovia M, Llewellyn MS, Morocoima A, Urdaneta-Morales S, et al. (2012) Geographical Distribution of Trypanosoma cruzi Genotypes in Venezuela. PLoS Negl Trop Dis 6(6): e1707.
Lauthier JJ, Tomasini N, Barnabé C, Rumi MM, D'Amato AM, Ragone PG, Yeo M, Lewis MD, Llewellyn MS, Basombrío MA, Miles MA, Tibayrenc M, Diosque P. (2012) Candidate targets for Multilocus Sequence Typing of Trypanosoma cruzi: validation using parasite stocks from the Chaco Region and a set of reference strains. Infect Genet Evol. 12(2):350-8
Messenger LA, Llewellyn MS, Bhattacharyya T, Franzén O, Lewis MD, et al. (2012) Multiple Mitochondrial Introgression Events and Heteroplasmy in Trypanosoma cruzi Revealed by Maxicircle MLST and Next Generation Sequencing. PLoS Negl Trop Dis 6(4): e1584 doi:10.1371/journal.pntd.0001584
Zingales B, Miles MA, Campbell DA, Tibayrenc M, Macedo AM, Teixeira MM, Schijman AG, Llewellyn MS, Lages-Silva E, Machado CR, Andrade SG, Sturm NR.(2012) The revised Trypanosoma cruzi subspecific nomenclature: rationale, epidemiological relevance and research applications. Infect Genet Evol.12:240-53
Llewellyn, M.S.,Rivett-Carnac, J.B., Fitzpatrick, S., Lewis, M. D., Yeo, M., Gaunt M.W., Miles M.A., Extraordinary Trypanosoma cruzi diversity within single mammalian reservoir hosts implies a mechanism of diversifying selection. Int J Parasitol. 41(6):609-14.
Franzen, O. Ochaya, S. Sherwood, E. Lewis, M. Llewellyn, M. 2011Miles, M. Andersson, B. 2011 Shotgun Sequencing Analysis of Trypanosoma cruzi I Sylvio X10/1 and comparison with T. cruzi VI CL Brener. PLoS Negl Trop Dis 5(3):e984
Yoshida, N. Tyler K.M., Llewellyn, M.S.2011Invasion mechanisms among emerging food borne protozoan parasites. Trends in Parasitology. 27(10):459-66
Yeo M, Mauricio IL, Messenger LA, Lewis MD, Llewellyn MS, Nidia Acosta1,Tapan Bhattacharyya, Patricio Diosque, Hernan J. Carrasco, Michael A. Miles (2011) Multilocus Sequence Typing (MLST) for Lineage Assignment and High Resolution Diversity Studies in Trypanosoma cruzi. PLoS Negl Trop Dis 5(6): e1049.
Ocaña-Mayorga S, Llewellyn MS, Costales JA, Miles MA, Grijalva MJ, 2010 Sex, Subdivision, and Domestic Dispersal of Trypanosoma cruzi Lineage I in Southern Ecuador. PLoS Negl Trop Dis 4(12): e915.
Hamilton, P.B., Lewis, M.D., Cruickshank, C., Gaunt, M.W., Yeo, M., Llewellyn, M.S.,Valente, S.A., Maia da Silva, F., Stevens, J.R., Miles, M.A., Teixeira, M.M., 2010 Identification and lineage genotyping of South American trypanosomes using fluorescent fragment length barcoding. Infect Genet Evol 11, 44-51.
Bhattacharyya, T., Brooks, J., Yeo, M., Carrasco, H.J., Lewis, M.D., Llewellyn, M.S., Miles, M.A., 2010 Analysis of molecular diversity of the Trypanosoma cruzi trypomastigote small surface antigen reveals novel epitopes, evidence of positive selection and potential implications for lineage-specific serology. Int J Parasitol. 40(8):921-8
Llewellyn, M.S.,Miles, M.A., Carrasco, H.J., Lewis, M.D., Yeo, M., Vargas, J., Torrico, F., Diosque, P., Valente, V., Valente, S.A., Gaunt, M.W., 2009. Genome-scale multilocus microsatellite typing of Trypanosoma cruzi discrete typing unit I reveals phylogeographic structure and specific genotypes linked to human infection. PLoS Pathog 5, e1000410.
Llewellyn, M.S., Lewis, M.D., Acosta, N., Yeo, M., Carrasco, H.J., Segovia, M., Vargas, J., Torrico, F., Miles, M.A., Gaunt, M.W., 2009. Trypanosoma cruzi IIc: Phylogenetic and Phylogeographic Insights from Sequence and Microsatellite Analysis and Potential Impact on Emergent Chagas Disease. PLoS Negl Trop Dis 3, e510.
Lewis, M.D., Llewellyn, M.S., Gaunt, M.W., Yeo, M., Carrasco, H.J., Miles, M.A., 2009. Flow cytometric analysis and microsatellite genotyping reveal extensive DNA content variation in Trypanosoma cruzi populations and expose contrasts between natural and experimental hybrids. Int J Parasitol 39, 1305-1317.
Lewis, M.D., Ma, J., Yeo, M., Carrasco, H.J., Llewellyn, M.S., Miles, M.A., 2009. Genotyping of Trypanosoma cruzi: systematic selection of assays allowing rapid and accurate discrimination of all known lineages. Am J Trop Med Hyg 81, 1041-1049.
Marcili, A., Lima, L., Valente, V.C., Valente, S.A., Batista, J.S., Junqueira, A.C., Souza, A.I., da Rosa, J.A., Campaner, M., Lewis, M.D., Llewellyn, M.S., Miles, M.A., Teixeira, M.M., 2009. Comparative phylogeography of Trypanosoma cruzi TcIIc: New hosts, association with terrestrial ecotopes, and spatial clustering. Infect Genet Evol. 9(6):1265-74
Miles, M.A., Llewellyn, M.S., Lewis, M.D., Yeo, M., Baleela, R., Fitzpatrick, S., Gaunt, M.W., Mauricio, I.L., 2009. The molecular epidemiology and phylogeography of Trypanosoma cruzi and parallel research on Leishmania: looking back and to the future. Parasitology 136, 1509-1528.
Yeo, M., Acosta, N., Llewellyn, M.,Sanchez, H., Adamson, S., Miles, G.A., Lopez, E., Gonzalez, N., Patterson, J.S., Gaunt, M.W., Arias, A.R., Miles, M.A., 2005. Origins of Chagas disease: Didelphis species are natural hosts of Trypanosoma cruzi I and armadillos hosts of Trypanosoma cruzi II, including hybrids. Int J Parasitol 35, 225-233.
Yeo, M., Lewis, M.D., Carrasco, H.J., Acosta, N., Llewellyn, M., da Silva Valente, S.A., de Costa Valente, V., de Arias, A.R., Miles, M.A., 2006. Resolution of multiclonal infections of Trypanosoma cruzi from naturally infected triatomine bugs and from experimentally infected mice by direct plating on a sensitive solid medium. Int J Parasitol 37, 111-120
.
Kassen, R., Llewellyn, M.,Rainey, P.B., 2004. Ecological constraints on diversification in a model adaptive radiation. Nature 431, 984-988.
