Past Research Projects
 Enhancing freshwater ecosystem biomonitoring: defining and testing an ecometagenetic DNA identification framework.
Enhancing freshwater ecosystem biomonitoring: defining and testing an ecometagenetic DNA identification framework.
Biological indicators are used to estimate the state of the environment and the Environment Agency mandatorily monitors the health of freshwater ecosystems throughout the UK. However, traditional monitoring approaches suffer from the “identification bottleneck” problem (e.g. subjectivity, lack of resolution, labour-intensive traditional taxonomy approaches mismatched with ecosystem diversity). This KESS-funded PhD program, in collaboration with Iliana Bista and the Environment Agency, will populate a DNA reference library for up to 200 macroinvertebrate indicator species and test and test the efficacy of the use of second generation sequencing approaches in enhancing freshwater biomonitoring programs. Chironomid image ©entomart.
Understanding the microbial mechanisms underpinning carbon release from peatland ecosystems.
In their natural waterlogged state, peatlands constitute the most important, long-term, terrestrial organic carbon store on our planet. However, it has been shown that under drought conditions the stored carbon is released as atmospheric gaseous CO2 and dissolved organic carbon (DOC) into surrounding waterways, contributing to global warming and the net loss of carbon from peatland ecosystems. Here, funded by the HPC Wales/Fujitsu partnership and partnered with Welsh Water, we will use “metagenetic” and shotgun “metagenomic” approaches to identify both the composition and functional diversity of the microbial communities  responsible for drought-driven carbon loss from a range of peatland ecosystems. By understanding the mechanisms associated with carbon loss, we will be able to explore approaches aimed at regulation of key microbial drivers of carbon loss through the geo-engineering of peatlands to regain carbon. Such a multidisciplinary project demands expertise in wetland biogeochemistry and metagenomics and is lead by Caitlin Potter, in collaboration with Prof. Chris Freeman, Prof. Peter Golyshin, Dr. Nathalie Fenner and Dr. James McDonald.
responsible for drought-driven carbon loss from a range of peatland ecosystems. By understanding the mechanisms associated with carbon loss, we will be able to explore approaches aimed at regulation of key microbial drivers of carbon loss through the geo-engineering of peatlands to regain carbon. Such a multidisciplinary project demands expertise in wetland biogeochemistry and metagenomics and is lead by Caitlin Potter, in collaboration with Prof. Chris Freeman, Prof. Peter Golyshin, Dr. Nathalie Fenner and Dr. James McDonald.
Glastir Monitoring and Evaluation Programme (GMEP): Soil biodiversity

Mitogenomics and molecular identification of spiders
Spiders are among the oldest and most diverse groups of terrestrial organisms, with a current diversity of over 37,500 described species placed in 3,471 genera and 109 families. From an ecological point of view, spiders are an unequivocally important guild. Furthermore, spiders are model organisms in biochemical (silk proteins and venom), behavioural (especially sexual and web-building behaviours), ecological (foraging, predator-prey systems, integrated pest management), comparative development and speciation research (http://research.amnh.org/atol/files/).
Advancing mitogenomics via ultrasequencing: A case study in the Araneae. 
Since the late 1980s, molecular systematics has progressed via the analysis of increasingly larger numbers of genes and characters. At one end of the spectrum, phylogenies are derived from a moderate number of gene partitions, and at the other, genome sequencing and interrogation of expressed sequence tag (EST) libraries are leading to phylogenomic approaches. A compromise between these extremes lies in mitogenomic analyses (the comparative analyses of whole mitochondrial genomes) that have been shown capable of resolving evolutionary relationships among a large range of higher taxa. Although the availability of mitogenomic datasets are increasing, there are still logistical limitations regarding the chain-termination sequencing of c. 15,000 b.p. of sequence data for multiple taxa. A clear solution to this problem lies within ultrasequencing platforms. Despite the ecological and evolutionary significance of the Araneae, our current ability to address comparative phylogenetic hypotheses is severely limited by a lack of a robust phylogenetic framework for the group as a whole. In collaboration with Andy Briscoe, Dr. Sara Goodacre (Nottingham University), Prof. Greg Hurst (University of Liverpool), Dr. Miquel Arnedo (Universitat de Barcelona) and Dr. Susan Masta (Portland State University) this CoSyst-funded project will seek to augment ongoing US AToL endeavours by sequencing and analysing a large number of spider mitogenomes in order to create a robust, family-level phylogenetic framework.
 The macroecology of marine benthic meiofaunal diversity
The macroecology of marine benthic meiofaunal diversity
Marine benthic biodiversity is important for ecosystem functioning, sustainability and resilience, but the magnitude and composition of marine diversity at a range of spatial and taxonomic scales are undefined. Marine meiofaunal communities are dominated by the potentially hyperdiverse nematodes, accompanied by other small metazoans from 60% of animal phyla, therefore presenting a major fraction of global biodiversity. Despite the abundance and pivotal role in ecosystem functioning, a current estimate of global meiobenthic diversity remains a matter of conjecture. This knowledge gap is the result of the small size and the apparent morphological similarity of microbial metazoans that causes problems in the logistics and accuracy of species identification. Over the past few years myself and collaborators have contributed substantially to the development of a new field of eukaryotic biodiversity identification, that uses "second", or "next" generation sequencing approaches to simultaneously assess the richness of multiple phyla from the marine benthos. Empirical and perspective style papers have been published in Molecular Ecology, Nature Communications, Nucleic Acids Research and Trends in Ecology and Evolution.
Are marine nematodes hyperdiverse?
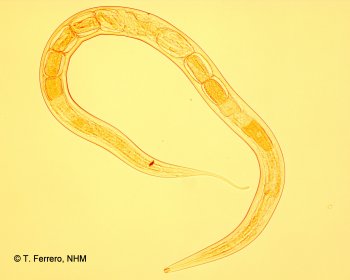 This NERC funded project used standard and novel molecular approaches (454 Roche 18S sequencing) and videocapture technology to estimate the actual molecular (and subsequent species) diversity present at different spatial scales throughout littoral communities of UK nematodes and other meiofauna and extrapolate this information to estimates of regional and global species richness. The utilization of second-generation sequencing to quantify molecular biodiversity represents a major advance towards identifying a crucial biological component of the earth's ecosystems and enable further hypotheses to be tested regarding the complex nature of 454 sequencing.
This NERC funded project used standard and novel molecular approaches (454 Roche 18S sequencing) and videocapture technology to estimate the actual molecular (and subsequent species) diversity present at different spatial scales throughout littoral communities of UK nematodes and other meiofauna and extrapolate this information to estimates of regional and global species richness. The utilization of second-generation sequencing to quantify molecular biodiversity represents a major advance towards identifying a crucial biological component of the earth's ecosystems and enable further hypotheses to be tested regarding the complex nature of 454 sequencing.
Sequencing the meiofaunal metagenome of the fresh/saltwater interface.
 Estuaries are key transitional habitats that are significantly affected by local and global anthropogenic activities. They are typically considered to be low diversity systems; a taxonomically biased inference drawn from the low alpha diversity of macrofauna. In contrast, meiofaunal diversity is substantial, with most estuaries estimated to be inhabited by approximately 200 species of nematodes, with numbers ranging from 106-108 animals per square metre, contributing to between 50-90% of the metazoan faunal species richness. In collaboration with The Environment Agency, The Thames Estuary Partnership and The Mersey Basin Campaign, this NERC funded project will link large-scale biodiversity appraisals (454 18S sequencing, videocapture and morphology) of environmental samples across ecological gradients with hydrodynamic models and macrofaunal processes.
Estuaries are key transitional habitats that are significantly affected by local and global anthropogenic activities. They are typically considered to be low diversity systems; a taxonomically biased inference drawn from the low alpha diversity of macrofauna. In contrast, meiofaunal diversity is substantial, with most estuaries estimated to be inhabited by approximately 200 species of nematodes, with numbers ranging from 106-108 animals per square metre, contributing to between 50-90% of the metazoan faunal species richness. In collaboration with The Environment Agency, The Thames Estuary Partnership and The Mersey Basin Campaign, this NERC funded project will link large-scale biodiversity appraisals (454 18S sequencing, videocapture and morphology) of environmental samples across ecological gradients with hydrodynamic models and macrofaunal processes.
Understanding the functional genomic mechanisms underpinning endocrine disruption in the ecological sentinel, Nucella lapillus
Man-made chemicals that disrupt endocrine function (endocrine dispuptors) contaminate all environments, producing a range of undesirable effects across a wide range of taxa and can cause abnormal development of male and female traits. 
